Organismi marini: genomica, sviluppo ed evoluzione
Dipartimenti coinvolti:
Sezione di Ricerca Scientifica "Biologia ed Evoluzione degli Organismi Marini" (BEOM) - leader
Sezione di Servizio e Ricerca Tecnologica "Infrastrutture di Ricerca per le Risorse Biologiche Marine" (RIMAR)
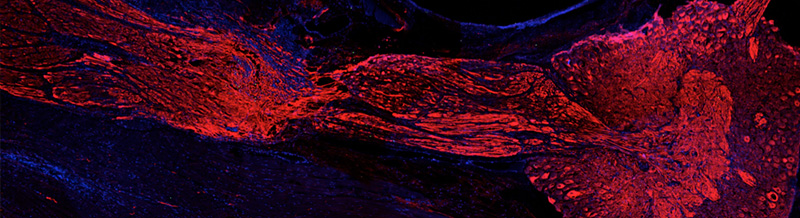
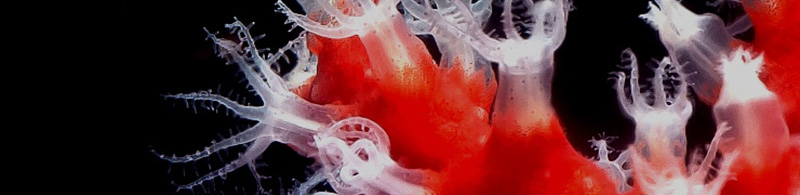
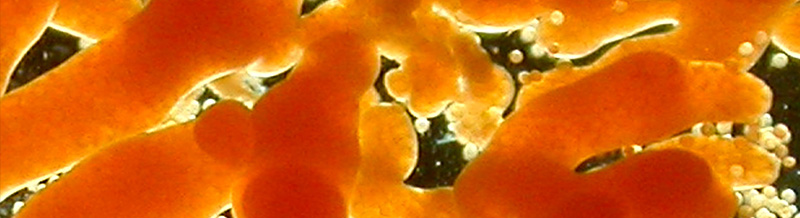
La SZN è riconosciuta internazionalmente per le competenze ed i contributi dati alla comprensione della biologia di diversi organismi marini che rappresentano modelli di studio ideali in diversi ambiti quali la biologia dello sviluppo, la riproduzione e l’evoluzione, fino alla sperimentazione pre-clinica. In continuità con la forte tradizione di studi su organismi marini, e trovandosi in una posizione ideale per le capacità di accesso a diversi ambienti/habitat marini del Mediterraneo, ben conosciuti perché oggetto di studio nell’ambito di altri progetti istituzionali, la SZN è impegnata ad identificare un gruppo più esteso di organismi marini da poter proporre come modello emergente per studi multidisciplinari che coprano diversi aspetti della ricerca di base ed applicativa.
La ricerca biologica si è basata per lungo tempo sullo studio di un limitato numero di "organismi modello" (e.g. Escherichia coli, lievito, Drosophila, rana, topo, Arabidopsis, Caenorhabditis elegans e pochissime altre specie). Tali organismi sono stati selezionati perché facilmente utilizzabili nelle ricerche di laboratorio e adatti per lo studio di una serie di studi biologici. La varietà di organismi utilizzati come "modelli" nella ricerca è attualmente oggetto di una massiccia espansione, grazie alla riduzione del tempo e dei costi del sequenziamento dei genomi e alla disponibilità di tecniche per alterare selettivamente i pattern di espressione dei geni. Inoltre, sempre più biologi espandono i loro interessi da quello puramente meccanicistico all’integrazione di tematiche evolutive. L’introduzione di nuove specie "al laboratorio" apre nuove vie di ricerca e permette la comparazione, l’avanzamento e l’espansione della nostra comprensione di processi biologici.
Gli organismi marini rappresentano quindi una importante fonte di nuovi "organismi modello". La SZN si pone l’obiettivo di creare un nuovo catalogo, diversificato per tipologia di utilizzo, con una nuova generazione di specie potenzialmente utili per estendere la ricerca scientifica in nuove direzioni.
Il progetto "Organismi marini: genomica, sviluppo ed evoluzione" coordinato dalla Sezione di Ricerca "Biologia ed Evoluzione degli Organismi Marini" della SZN, si articola in tre obiettivi:
- Sequenziamento del genoma di 100 organismi "modello" del Mar Mediterraneo.
- Studio dell’origine ed evoluzione dei meccanismi di sviluppo nei deuterostomi.
- Studio della plasticità biologica degli organismi marini.
Osservatorio Marino: Biodiversità e Funzionamento degli Ecosistemi
Dipartimenti coinvolti:
Sezione di Servizio e Ricerca Tecnologica "Infrastrutture di Ricerca per le Risorse Biologiche Marine" (RIMAR) – leader
Sezione di Ricerca Scientifica "Ecologia Marina Integrata" (EMI)
Il Mar Mediterraneo è il più grande mare semi-chiuso del mondo, comprende 29 paesi e territori di Europa, Africa e Medio Oriente. In una delle regioni più popolate del globo, la regione mediterranea è considerata come un hotspot di biodiversità. Nonostante rappresenti meno dell'1% degli oceani del mondo, il Mediterraneo conta oltre il 7,5% di tutte le specie marine conosciute, con un'alta percentuale di specie endemiche. Minacciato da inquinamento, urbanizzazione, turismo e pesca eccessiva, il Mar Mediterraneo ospita numerose specie in pericolo (e.g. foca monaca, tartarughe marine, cetacei). Nonostante non sia stato ancora riportato alcun evento di estinzione di specie autoctone, sono stati registrati improvvisi cali di abbondanza ed estirpazioni locali, in concomitanza con l’invasione di specie aliene. In questo contesto, la Lista Rossa del Mediterraneo sviluppata dal Centro per la Cooperazione Mediterranea dell'IUCN mira a valutare lo stato di conservazione della regione mediterranea.
La SZN ha avviato la realizzazione di un Osservatorio Marino integrato, che possa per la prima volta in Italia mettere insieme le osservazioni ambientali ed oceanografiche con le componenti biologiche ed ecologiche. L’obiettivo è quello di integrare in un contesto sinergico lo studio della biodiversità e del funzionamento degli ecosistemi marini, contribuendo alla loro gestione eco-sostenibile. L’osservatorio è rivolto al Mar Tirreno, con una attenzione particolare al Golfo di Napoli. Attraverso osservazioni sul campo si mira ad ottenere una mappatura della biodiversità nello spazio e nel tempo, correlandola con le proprietà chimiche e fisiche dell'acqua, le interazioni biotiche, l’identificazione di nuove specie, comprese le specie invasive, e lo sviluppo di strumenti di monitoraggio automatici.
La definizione quantitativa e qualitativa della biodiversità rappresenta un contributo fondamentale alla corretta gestione ambientale, al raggiungimento del buono stato ecologico (GES, sensu Marine Strategy Framework Directive MSFD Direttiva Quadro sulla Strategia Marina) e della buona qualità ambientale, che a loro volta, sono essenziali per il benessere umano e lo sfruttamento sostenibile delle risorse marine. L'acquisizione di queste conoscenze si inserisce nelle tematiche del Piano Nazionale della Ricerca (Blue growth) e Horizon2020 "Tutela dell'ambiente marino come fonte di sostentamento, l'energia e le biotecnologie e per la scelta degli strumenti necessari per le decisioni dei responsabili politici".
Questo ambizioso obiettivo si deve attuare attraverso un approccio “end-to-end” sia per quanto riguarda il livello di organizzazione biologica (dalle cellule agli ecosistemi), sia per quanto concerne le integrazioni di dati di diversità strutturale e funzionale, la capacità di utilizzare tecnologie avanzate e sofisticate quali i sistemi di “osservazione biologica” in situ, sia per quanto concerne il loro campionamento. Alle molteplici attività di campionamento vengono affiancati studi in mesocosmi sperimentali, in laboratorio ed analisi funzionali che mirano a definire i meccanismi alla base del funzionamento degli organismi e degli ecosistemi marini oggetto di studio.
L’utilizzo di un approccio modellistico permetterà infine integrare le diverse scale di osservazione, dalla cellula al sistema mare su scala regionale per trarne informazioni utili sia alla gestione che alla conservazione delle risorse marine.
Questo progetto si attua attraverso il perseguimento dei seguenti obiettivi:
- Osservatorio marino del Tirreno: approccio multidisciplinare allo studio a lungo termine della biologia ed ecologia marina (Long -Term Ecological Research).
- Biodiversità e Funzionamento degli ecosistemi marini.
- Conservazione Biologica Marina.
Giuseppina Barsacchi
Dipartimento di Biologia
Università di Pisa
Ferdinando Boero
Dipartimento di Scienze e Tecnologie Biologiche e Ambientali
Università del Salento, Lecce
Gary G. Borisy
Director Marine Biological Laboratory (MBL)
Woods Hole, Mass. Usa
Peter Bukill
Director of the Sir Alister Hardy Foundation for Ocean Sciences and professor of Ocean Science, University of Plymouth, Plymoth, UK
Rita R. Colwell
Professor at University of Maryland College Park and at Johns Hopkins University Bloomberg School of Public Health Environmental Health Sciences, Center for Water and Health
Baltimore, MD, USA
Paul K. Dayton
Integrative Oceanography Division
Scripps Institution of Oceanography
University of California
San Dego, La jolla, CA, USA
Carlos Duarte
Director Instituto Mediterraneo de Estudio Avanzados
Mallorca, Illes Balears, SPAIN
Aldo Fasolo
Dipartimento di Biologia Animale e dell’uomo
Università di Torino
Richard Timothy Hunt – Premio Nobel per la Medicina 2001
London Research Institute, Clare Hall Laboratories
London, UK
Thomas Kiørboe
Professor, Technical University of Denmark
Danish Institute for Fisheries Research
Afd. for Havøkologi og Akvakultur, Kavalergården 6
Charlottenlund, DENMARK
Iain Mattaj
Director General European Molecular Biology Laboratory EMBL
Heidelberg, GERMANY
Christiane Nüsslein-Volhard - Premio Nobel per la Medicina 1995
Director of Dept. of Genetics
Max Planck Institute for Developmental Biology
Tübingen, GERMANY
Noriyuki Satoh
Department of Zoology, Kyoto University
Kyoto, JAPAN
Torsten Wiesel – Premio Nobel per la Medicina 1981
Vincent and Brooke Astor Professor Emeritus and President Emeritus
The Rockefeller University, Laboratory of Neurobiology
New York, NY, USA
Contributo della Biologia Marina e delle Blue biotechnologies alla "Blue Growth"
Dipartimenti coinvolti:
Sezione di Ricerca Scientifica "Ecologia Marina Integrata" (EMI) – leader
Sezione di Ricerca Scientifica "Biologia ed Evoluzione degli Organismi Marini" (BEOM)
Sezione di Servizio e Ricerca Tecnologica "Infrastrutture di Ricerca per le Risorse Biologiche Marine" (RIMAR)
La Biologia marina, la scoperta di nuove forme di vita, la conoscenza dei loro adattamenti e della loro ecologia costituiscono la base di ricerca pura su cui si basano le più importanti scoperte scientifiche in questo settore. È su questo terreno che la Stazione Zoologica sviluppa il proprio approccio alle biotecnologie marine (ovvero l'applicazione di tecniche avanzate e conoscenze innovative per sviluppare prodotti biologici e altri fattori benefici per gli esseri umani).
Le Blue Biotechnologies sono in costante crescita in Europa e nel panorama internazionale, e contribuiranno sempre di più a plasmare il futuro delle nostre società. Le biotecnologie marine, che implicano lo studio e la conoscenza delle risorse biologiche marine, stanno rapidamente diventando una componente importante del settore Blue Growth.
Tra gli obiettivi di Horizon 2020 nel settore della crescita blu troviamo:
"applicare, in modo etico e sostenibile, strumenti avanzati per fornire un contributo significativo alla soluzione delle principali sfide sociali nei settori della sicurezza alimentare ed energetica, lo sviluppo di nuovi farmaci e trattamenti per la salute umana e animale, materiali industriali e processi e l’uso e gestione sostenibile dei mari e degli oceani"
(Ocean Research in Horizon 2020: the Blue Growth Potential)
Questo Progetto della SZN ha l'ambizione di affrontare diverse linee chiave della ricerca che sono essenziali nel settore delle "biotecnologie blu". Esso copre molti campi di ricerca e si attua attraverso lo svolgimento dei seguenti obiettivi:
- Biodiversità marina: ecologia degli organismi marini ed identificazione di specie di interesse biotecnologico.
- Potenziale biotecnologico di organismi marini in campo farmaceutico.
- Potenziale biotecnologico degli organismi marini in campo nutraceutico, cosmetico e ambientale.
Giuseppina Barsacchi
Dipartimento di Biologia
Università di Pisa
via G. Carducci, 13
56010 Ghezzano (Pisa) - Italia
 Peter Burkill
Peter Burkill
Marine Institute
Plymouth University
Drake Circus
Plymouth PL4 8AA - United Kingdom
Rita R. Colwell
Center for Bioinformatics and Computational Biology
University of Maryland
296 Biomolecular Sciences Bldg.
Room 3103
College Park, MD 20742 - USA
Roberto Danovaro
Dipartimento Scienze del Mare
Università Politecnica delle Marche
Via Brecce Bianche
60131 Ancona - Italia
Aldo Fasolo
Dipartimento di Biologia Animale e dell’Uomo
Università di Torino
Via Accademia Albertina, 13
10123 Torino - Italia
Bernard Kloareg
Station Biologique Roscoff
Place Georges Teissier - BP74
29682 Roscoff Cedex - France
Noriyuki Satoh
Marine Genomics Unit
Okinawa Institute of Science and Technology
1919-1 Tancha, Onna-son
Kunigami-gun
904-0495 - Japan